


Find out how you can use REANA to describe, run, preserve and reuse your analyses.
User GuideInstall and manage the REANA reusable analysis platform on your own compute cloud.
Administrator GuideUnderstand REANA source code, adapt it to your needs, contribute changes back.
Developer Guide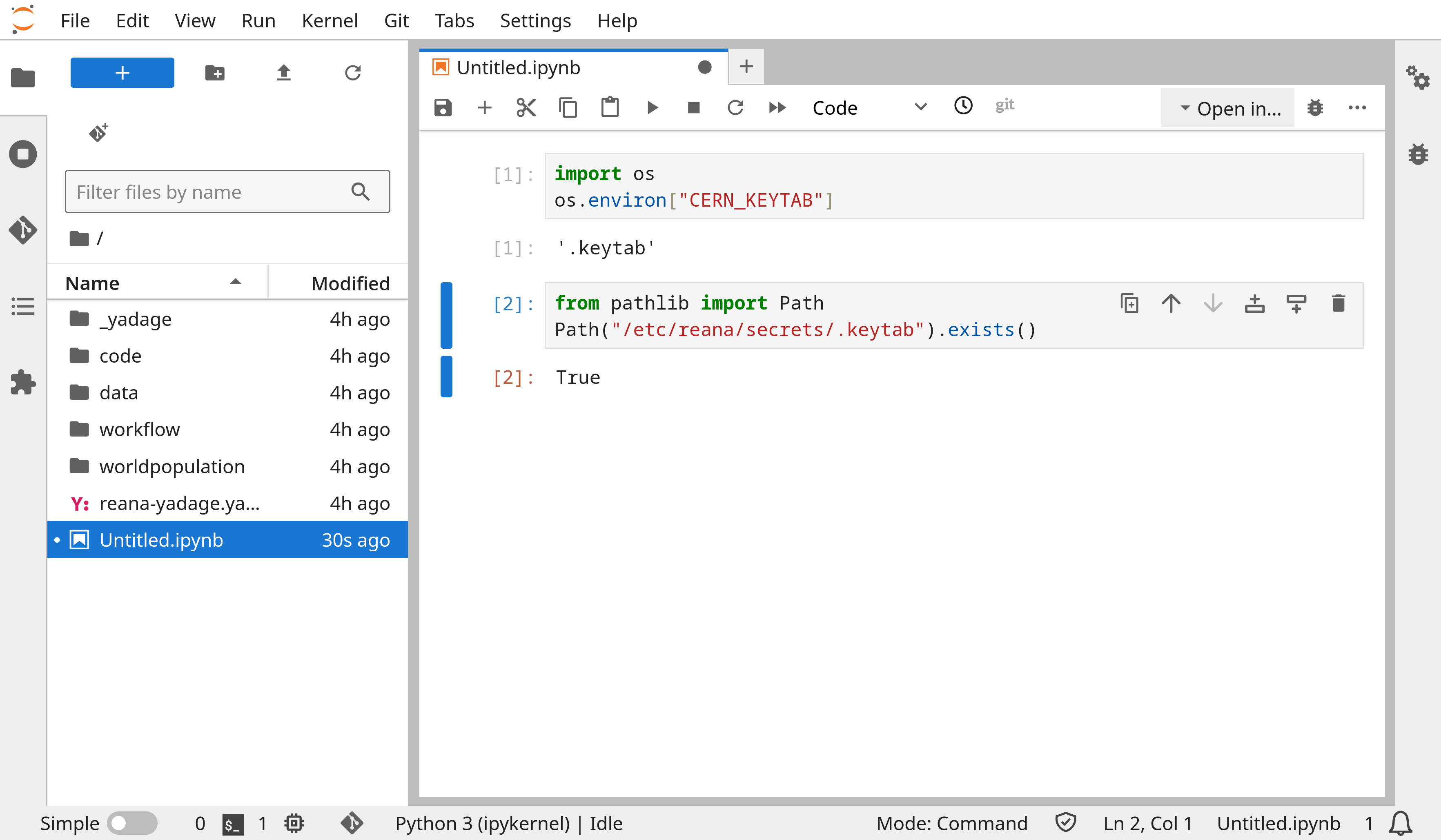
Adds support for user secrets in Jupyter notebook sessions.
Introduces support for Compute4PUNCH infrastructure.
Enhances HTCondor compute backend job dispatch.
Strengthens the security of the platform.
| March 24th-28th 2025: Alp and Tibor participate at the HSF/IRIS-HEP Analysis Reproducibility (Virtual) trainig event. [event] [slides] | |
| March 17th-18th 2025: Enrique and Jose participate at the Kick-off workshop of the EOSC Federation. [event] | |
| February 25th-28th 2025: Giovanni presents CERN platforms for scientific analyses (including REANA) at the (Re)interpretation of the LHC results for new physics workshop. [event] [slides] | |
| February 17-18th 2025: Tibor presents REANA: status and plans at the CMS Physics Days event focused on Beyond the Standard Model Combinations. [slides] | |
| February 14th 2025: Alp presents Supporting Dask workloads in the REANA reproducible analysis platform at the IRIS-HEP Analysis Grand Challenge Demo Day 7. [event] [slides] | |
| November 24th-29th 2024: Tibor runs a reproducible workflow training at the EURO-LABS Advanced Training: Open Science and Data Management in Ebernburg, Germany. [event] [slides] | |
| November 4th-8th 2024: Marco and Tibor run a reproducible workflow training at the 1st CERN School of Computing on IT Services in Ferney-Voltaire, France. [event] [slides] [exercises] | |
| October 19th-25th 2024: Marco participates at the Conference on Computing in High Energy and Nuclear Physics (CHEP 2024) in Krakow, Poland. [event] [slides] | |
| October 7th 2024: Jelizaveta presents Improving latency and scalability of the user runtime job log collecting and exposure in REANA at the IRIS-HEP Fellows Final Presentations event. [event] [slides] | |
| October 2nd-3rd 2024: Marco and Tibor participate at the 4th DPHEP Collaboration Workshop in Geneva, Switzerland. [event] [slides] | |
| September 17th-18th 2024: Tibor participates at the 2nd Next Generation Environment for Interoperable Data Analysis Expert Workshop in Dortmund, Germany. [event] [slides] | |
| August 28th 2024: Andrii presents Analysis Grand Challenge on REANA at the IRIS-HEP Gap Year Final Presentations event. [event] [slides] | |
| June 20th 2024: Tibor presents REANA at the Swiss Reproducibility Network comuptational reproducibility seminar. [series] [slides] | |
| June 18th-20th 2024: Tibor participates at the Analysis Facilities Workshop in Garching, Germany. [event] [slides] | |
| April 3rd-5th 2024: Andrii, Marco and Tibor participate at the Workshop on workflow languages for HEP analysis in Geneva, Switzerland. [event] [slides] | |
| April 2nd 2024: REANA featured in the Analysis Facilities White Paper. [paper] | |
| February 26th-March 1st 2024: Tibor presents at the HSF Training on Analysis Pipelines. [event] [slides] |
Introduce role-based authorisation control models allowing to share workflows with colleagues.
Allow users to dynamically create Dask cluster necessary for their workflows.
Introduce new command-line client written in Go to improve performance and ease of use. Stabilise the REST API versioning.